- Infectious Diseases of Livestock
- Part 2
- Porcine epidemic diarrhoea
- GENERAL INTRODUCTION: PARAMYXOVIRIDAE AND PNEUMOVIRIDAE
- Rinderpest
- Peste des petits ruminants
- Parainfluenza type 3 infection
- Bovine respiratory syncytial virus infection
- Hendra virus infection
- Paramyxovirus-induced reproductive failure and congenital defects in pigs
- Nipah virus disease
- GENERAL INTRODUCTION: CALICIVIRIDAE AND ASTROVIRIDAE
- Vesicular exanthema
- Enteric caliciviruses of pigs and cattle
- GENERAL INTRODUCTION: RETROVIRIDAE
- Enzootic bovine leukosis
- Jaagsiekte
- Visna-maedi
- Caprine arthritis-encephalitis
- Equine infectious anaemia
- GENERAL INTRODUCTION: PAPILLOMAVIRIDAE
- Papillomavirus infection of ruminants
- Papillomavirus infection of equids
- GENERAL INTRODUCTION: ORTHOMYXOVIRIDAE
- Equine influenza
- Swine influenza
- GENERAL INTRODUCTION: CORONAVIRIDAE
- Porcine transmissible gastroenteritis
- Porcine respiratory coronavirus infection
- Porcine epidemic diarrhoea
- Porcine haemagglutinating encephalomyelitis virus infection
- Porcine deltacoronavirus infection
- Bovine coronavirus infection
- Ovine coronavirus infection
- Equine coronavirus infection
- GENERAL INTRODUCTION: PARVOVIRIDAE
- Porcine parvovirus infection
- Bovine parvovirus infection
- GENERAL INTRODUCTION: ADENOVIRIDAE
- Adenovirus infections
- GENERAL INTRODUCTION: HERPESVIRIDAE
- Equid herpesvirus 1 and equid herpesvirus 4 infections
- Equid gammaherpesvirus 2 and equid gammaherpesvirus 5 infections
- Equine coital exanthema
- Infectious bovine rhinotracheitis/infectious pustular vulvovaginitis and infectious pustular balanoposthitis
- Bovine alphaherpesvirus 2 infections
- Malignant catarrhal fever
- Pseudorabies
- Suid herpesvirus 2 infection
- GENERAL INTRODUCTION: ARTERIVIRIDAE
- Equine viral arteritis
- Porcine reproductive and respiratory syndrome
- GENERAL INTRODUCTION: FLAVIVIRIDAE
- Bovine viral diarrhoea and mucosal disease
- Border disease
- Hog cholera
- Wesselsbron disease
- Louping ill
- West nile virus infection
- GENERAL INTRODUCTION: TOGAVIRIDAE
- Equine encephalitides caused by alphaviruses in the Western Hemisphere
- Old World alphavirus infections in animals
- Getah virus infection
- GENERAL INTRODUCTION: BUNYAVIRIDAE
- Diseases caused by Akabane and related Simbu-group viruses
- Rift Valley fever
- Nairobi sheep disease
- Crimean-Congo haemorrhagic fever
- GENERAL INTRODUCTION: ASFARVIRIDAE
- African swine fever
- GENERAL INTRODUCTION: RHABDOVIRIDAE
- Rabies
- Bovine ephemeral fever
- Vesicular stomatitis and other vesiculovirus infections
- GENERAL INTRODUCTION: REOVIRIDAE
- Bluetongue
- Ibaraki disease in cattle
- Epizootic haemorrhagic disease
- African horse sickness
- Equine encephalosis
- Palyam serogroup orbivirus infections
- Rotavirus infections
- GENERAL INTRODUCTION: POXVIRIDAE
- Lumpy skin disease
- Sheeppox and goatpox
- Orf
- Ulcerative dermatosis
- Bovine papular stomatitis
- Pseudocowpox
- Swinepox
- Cowpox
- Horsepox
- Camelpox
- Buffalopox
- GENERAL INTRODUCTION: PICORNAVIRIDAE
- Teschen, Talfan and reproductive diseases caused by porcine enteroviruses
- Encephalomyocarditis virus infection
- Swine vesicular disease
- Equine picornavirus infection
- Bovine rhinovirus infection
- Foot-and-mouth disease
- GENERAL INTRODUCTION: BORNAVIRIDAE
- Borna disease
- GENERAL INTRODUCTION: CIRCOVIRIDAE AND ANELLOVIRIDAE
- Post-weaning multi-systemic wasting syndrome in swine
- GENERAL INTRODUCTION: PRION DISEASES
- Scrapie
- Bovine spongiform encephalopathy
- Transmissible spongiform encephalopathies related to bovine spongiform encephalopathy in other domestic and captive wild species
Porcine epidemic diarrhoea
This content is distributed under the following licence: Attribution-NonCommercial CC BY-NC  View Creative Commons Licence details here
View Creative Commons Licence details here
Porcine epidemic diarrhoea
Previous Authors: M B PENSAERT
Current Authors:
Q WANG - Associate Professor, PhD, Ohio State University, 1680 Madison Avenue, Wooster, Ohio, OH 44691, USA
L FENG - Head of Swine Digestive System Infectious Diseases, Head of Scientific Research Management and International Cooperation State Key Laboratory of Veterinary Biotechnology, Harbin Veterinary Research Institute of CAAS, No. 678 Haping Street, Xiangfang District, Harbin 150001, Heilongjiang, China
L J SAIF - Professor, MS, PhD, Food Animal Health Research Program, CFAES and CVM, OARDC, Ohio State University, 1680 Madison Ave, Wooster, Ohio, OH 44691, USA
Introduction
Infection with porcine epidemic diarrhoea virus (PEDV), a coronavirus, causes syndromes associated with fatal diarrhoea in neonatal pigs and transient diarrhoea in weaner, fattening and adult pigs.
Outbreaks of diarrhoea similar to transmissible gastroenteritis (TGE) but without the involvement of suckling piglets and caused by a coronavirus distinct from the TGE virus (TGEV), were first reported in England in 1971.71 Porcine epidemic diarrhoea virus was first isolated in 1978 and was found to be serologically unrelated to the other porcine coronaviruses.50, 51 It was subsequently also associated with diarrhoea in neonatal piglets.51, 71 Later, severe PEDV-induced outbreaks of diarrhoea in pigs of all ages were reported in Asia.30, 34, 60 In Europe in the 1970s-1980s, PEDV caused widespread epidemics with severe losses in suckling pigs. Then, PEDV became endemic in weaned and feeder pigs, with epidemics occurring occasionally.52 In Asia, epidemics with important losses in suckling pigs were first reported in the 1980s and continued to occur in the 1990s and 2000s. Before October 2010, PEDV infections were endemic in Chinese pig populations, but there were no reports of large-scale outbreaks.61 After that, highly virulent PEDV strains emerged in China, causing high mortality rates (up to 100 per cent) in neonatal piglets.62 In 2013, PEDV emerged for the first time in North America and caused large-scale outbreaks.59 By 2017, PEDV had been detected in many countries in Europe, Asia, and the Americas, but its presence in Australia and Africa has not been reported.
Aetiology
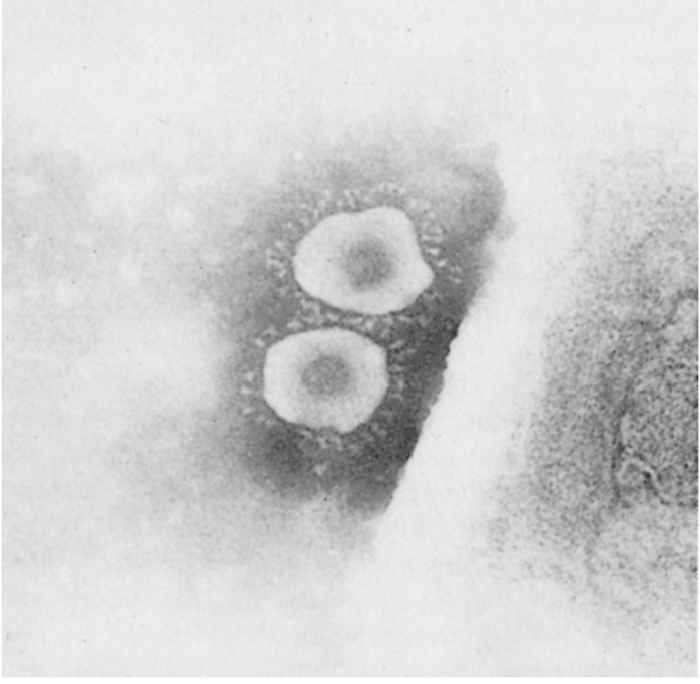
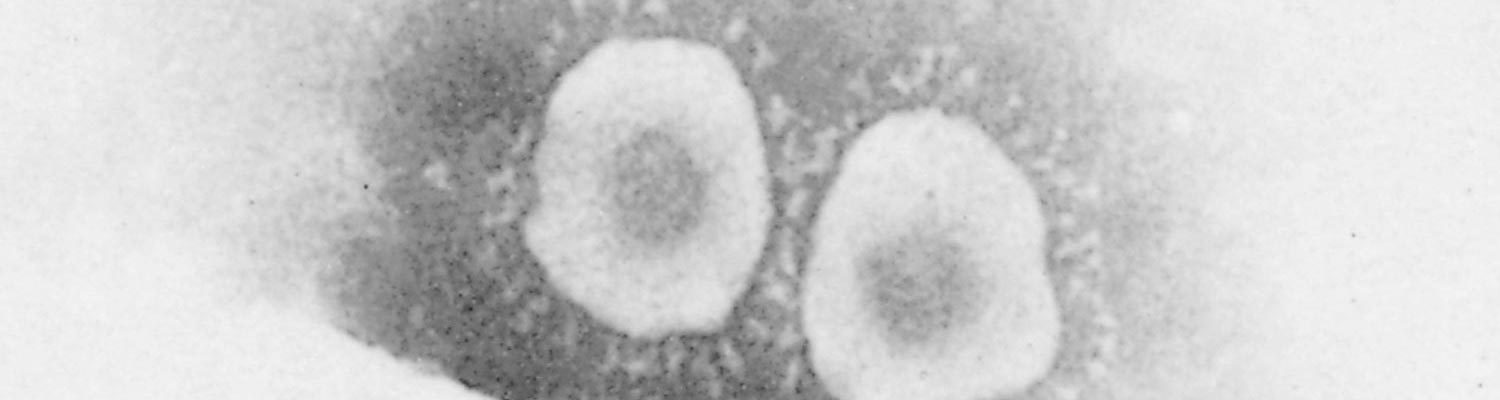
Morphologically, the virions of PEDV have the typical appearance of a coronavirus. They are nearly spherical or pleomorphic and have a mean overall diameter of 130 nm and a fringe of club-shaped projections with a mean length of 20 nm50 (Figure 1).
The virus has four structural proteins: a non-glycosylated RNA-binding nucleocapsid (N) protein, a small glycosylated membrane (M) protein, a small enveloped (E) protein, and a large glycosylated spike (S) protein that constitutes the surface projections.24, 57 The S protein is the receptor binding protein and contains viral neutralizing (VN) and non-VN epitopes.
Based on common antigenic determinants located on the nucleocapsid protein, PEDV was shown to be antigenically related to feline infectious peritonitis virus and TGEV.41, 72 Antigenic relationships to other members of the family Coronaviridae have not been found. On the basis of genomic analysis, PEDV is most closely related to bat alphacoronaviruses.29
There are no indications that different serotypes of PEDV exist.41 Genetically, PEDV strains can be divided into two major groups: classical PEDV strains and emerging PEDV strains.42, 66 Isolates from Europe and Asia that are genetically similar to the prototype CV777 strain originally isolated in Belgium are referred to as classical PEDVs. The highly virulent strains that emerged originally in China in 2010 and have spread to the Americas, Europe and other countries of Asia are referred to as emerging non-S INDEL PEDV. Recombinants between these two major groups of PEDV are called S INDEL strains and circulate in Asia, Europe and the Americas. Currently, no classical PEDV strains have been identified in the Americas. Other minor PEDV variants have been reported. Emerging non-S INDEL PEDV strains bearing large deletions (194-216 aa) in the N-terminal domain (NTD) of the S protein are designated as S1 NTD-del type of PEDV.21, 45, 63 Unlike the altered tissue tropism from enteric TGEV to respiratory for the TGEV variant, porcine respiratory coronavirus (PRCV), these PEDV strains have retained their enteric tropism, but with reduced virulence.28, 64 Furthermore, recombinant swine enteric coronaviruses (SeCoVs), i.e. recombinants between TGEV (backbone) and PEDV (S protein) have been reported in Europe.5, 14
The virus is sensitive to ether and chloroform. It loses its infectivity when heated at 60 °C for 30 minutes, but it is moderately stable at 50 °C. It is stable between pH 5 and 9 at 4 °C.
Porcine epidemic diarrhoea virus can be cultivated in Vero cells supplemented with trypsin in a serum-free medium. Not all Vero cell lines are equally susceptible. Cytopathic effects are characterised by the formation of syncytia.26, 45
Epidemiology
The pig is the only species that is known to be naturally infected by PEDV. Infection occurs by the faecal–oral route. Infected pigs can excrete infectious PEDV in faeces for about two weeks.16
Currently, classical PEDV strains are circulating in Asia and Europe, while emerging PEDV strains (non-S INDEL and S INDEL) are present in Asia, Europe and the Americas.42 The S1 NTD-del type strain was detected on pig farms in Japan and the USA (Wang, unpublished data) and most of these strains co-infected pigs with viruses possessing an intact S gene.21
Porcine epidemic diarrhoea virus causes up to 100 per cent mortality in neonatal piglets less than one week of age, and milder enteritis in older pigs. In Asian countries, PEDV still causes large outbreaks and substantial economic losses. In China, millions of piglets have died from PED since 2010 when highly virulent non-S INDEL PEDV strains emerged.67 In Japan, more than 490,000 pigs died due to PEDV during October 2013 to August 2015.21 In Thailand in 2007-2008, several outbreaks of severe PED were reported with Thai PEDV isolates in the same clade phylogenetically as the Chinese strain JS-2004-2 based on partial S sequence.55 Despite extensive usage of PEDV vaccines, PED was epidemic in South Korea during 2013-2014.15, 49 In Vietnam, PED was first reported in 2009 and has spread rapidly throughout the country causing huge economic losses.22 In the USA, about 10 per cent of the US pig population (7 million piglets) died during the first year (2013-2014) of PEDV outbreaks.42 The US strains identified during the initial outbreaks in the spring 2013 were non-S INDEL strains and closely related to Chinese PEDV strains (e.g. AH2012) reported in 2011-2012.29, 66 The first S INDEL strain was discovered in early 2014 in the US.69 However, a retrospective study found that S INDEL strains appeared in the US at roughly the same time as non-S INDEL strains.66 Moreover, these strains were found in diarrhoeic piglets in Japan, South Korea and Taiwan during late 2013 and early 2014.13, 43, 63 By summer 2017, PEDV had spread throughout the US and to other countries in North America, including Canada and Mexico.66 Currently PEDV outbreaks occur only sporadically in the USA. In Europe, outbreaks of diarrhoea due to PEDV are now endemic and mainly caused by emerging S INDEL PEDV strains in many countries (Austria, Belgium, France, Germany, Italy, The Netherlands, Portugal, Slovenia), except for an outbreak in the Ukraine caused by the emerging non-S INDEL PEDV.14
In four week -old pigs, infectious PEDV was shed in faeces for 14–16 days after infection.16 However, some pigs still shed PEDV RNA in faeces at 42 days after initial exposure, illustrating a discordance between prolonged detection of PEDV RNA in faeces and the transmission of infectious PEDV to susceptible pigs. No chronic or persistent infection of PEDV was reported. If there are several separate units on the farm, it may take several weeks for the virus to spread to all units.
After the disease has occurred on a breeding farm, the virus may either disappear or persist. The latter occurs mainly on farms containing large numbers of pigs born throughout the year, thus providing a regular source of susceptible animals. Alternatively, introduction of susceptible pigs onto farms before outbreak viruses have been eliminated enables the virus to persist. Porcine epidemic diarrhoea may then become a regular cause of diarrhoea which appears two to three weeks after weaning.8
Recently, PEDV was detected in 9.75 per cent of wild boars in South Korea,38 raising concerns that wild boars may play a role in maintenance and transmission of PEDV.
Pathogenesis
Porcine epidemic diarrhoea virus replicates in villous enterocytes in the small intestine and colon, causing cell necrosis and desquamation of infected cells in the small intestine, which leads to malabsorption of nutrients from the gut.19, 33 PEDV-infected cells were observed occasionally in the crypt area.42 The pathogenesis of classical PEDV infection in neonatal piglets is similar to that of TGEV infection, but differs in that in classical PEDV infection the events are somewhat slower and villous shortening is less pronounced. Therefore, the incubation period of PED caused by classical strains is longer, disease spread on farms slower, and the mortality lower (on average 50 per cent). The pathological features of classical PEDV infection in neonates in Asia are identical to those observed in Europe.
Variations in virulence of different PEDVs have been reported as indicated above. Emerging non-S INDEL PEDV strains gave rise to more virulent viruses than classical PEDV, emerging S INDEL PEDV strains, and the S1 NTD-del type of PEDV strains in neonatal pigs based on both field observations and experimentally infected pigs.11, 28, 33, 40, 44, 64 No direct comparison of pathogenicity has been reported for non-S INDEL PED and TGE viruses or the other three types of PEDV.
During the acute phase of PEDV infection, viral RNA was detected transiently in the serum of PEDV-infected suckling and weaned pigs.32, 33 Peak RNA levels in serum were about 4 log10 lower than RNA levels in faeces. Whether detection of viral RNA in serum represents infectious virus and a potential role in PEDV pathogenesis is unknown.
Porcine epidemic diarrhoea virus infection induces immunity similar to TGEV infection. Naturally infected sows are able to provide passive lactogenic immunity that protects suckling piglets from disease.37
Clinical signs
Porcine epidemic diarrhoea caused by different PEDV strains with high or low virulence may cause clinical signs that resemble those resulting from epidemic TGEV or PDCoV infections. When PEDV infects a population that is not fully susceptible, one of two different syndromes usually occurs. The first is diarrhoea in weaned grower and adult pigs, i.e. suckling piglets remain unaffected. For example, lactating sows may develop illness while their offspring remain healthy. The other syndrome, experienced when the disease first occurred in Europe, involves both neonatal and older pigs.
Fattening pigs can be severely affected and manifest abdominal pain and watery diarrhoea. In general, recovery occurs after seven to ten days. Mortality in the early stages of the disease may reach 1 to 2 per cent. In the syndrome where neonatal piglets are affected, diarrhoea and vomiting result in mortality due to dehydration in 50-100 per cent of the cohort. The acute disease is self-limiting as immunity is established two-three weeks after infection.20
Pathology
No gross pathognomic lesions are evident in animals that die during the acute stages of the disease. Microscopic lesions are characterized by necrosis and desquamation of enterocytes on the villi and villous shortening in the small intestine in piglets that die during the acute stage of diarrhoea.23, 54 These intestinal lesions are similar to those which occur in TGE and PDCoV infections. Microscopic lesions in the large intestine have not been described although PEDV infects < 30 per cent of colonic epithelial cells.42
Diagnosis and differential diagnosis
It is not possible to make an aetiological diagnosis of PED on clinical and pathological grounds alone. Acute PED outbreaks in which diarrhoea is observed in animals of all ages, including neonatal piglets, cannot be clinically differentiated from TGE or PDCoV. The laboratory diagnosis of PEDV infection was reviewed recently.20
An aetiological diagnosis can be made in the laboratory by detection of viral RNA and viral antigens in the tissues and faeces, and by the detection of antibodies in the serum and probably in oral fluid samples.4, 47
Vero cells are the most commonly used cell line for PEDV isolation; the success-rate for PEDV isolation is higher with intestinal contents/homogenates than with faeces.12, 26, 45 Immunofluorescence or immunohistochemistry on sections of the small intestine of pigs sacrificed during the acute stage of the disease are suitable for the diagnosis of PED.18 As this requires the sacrifice of an affected animal, it is used infrequently for the diagnosis of the disease in older pigs.
The detection of PEDV RNA is probably the most widely used laboratory diagnostic method for PED and for differentiation from other coronaviruses or viral pathogens causing diarrhoea. Reverse transcription (RT)-PCR and real-time RT-PCR (RT-qPCR) are sensitive, specific and rapid for the detection of viral RNA in clinical samples (faeces, serum, tissues, etc.). Because swine enteric coronaviruses (SeCoVs) are recombinants between TGEV (backbone) and PEDV (mainly S gene),5 only assays targeting both TGEV (any genes except for S gene) and PEDV (S gene) fragments are apt for differentiation of those viruses. Newer technology, such as the specific primer-independent next generation sequencing technology can be used to determine the viral genome sequences from faecal specimens. In situ hybridization can also be used to detect PEDV RNA in fixed tissues.
An antigen ELISA system using polyclonal or monoclonal antibodies can be used to detect PEDV in faecal material of older pigs. However, it is not often used for diagnostic purposes nowadays when more rapid and sensitive RT-PCR and RT-qPCR are available. This assay is more sensitive than direct detection of virions in the faeces by immune-electron microscopy, and is reliable if faeces are collected not later than two days after the onset of diarrhoea.7
For the serological diagnosis of the disease, paired acute and convalescent phase serum samples should be collected at an interval of two to four weeks for the virus neutralizing antibodies or IgG antibodies. A serological diagnosis is less feasible in pigs born of immune mothers after they have developed PEDV-induced diarrhoea shortly after weaning despite the presence of circulating maternal antibodies, as no clear-cut seroconversion may occur, but in theory one could assay for IgM antibodies indicative of a primary immune response to PEDV.
Serological diagnosis can be achieved by the use of either indirect immunofluorescence or immunofluorescence-blocking tests applied to PEDV-infected intestinal tissues or cell monolayers containing PEDV antigens70 Blocking ELISAs may also be performed using viral antigens derived from faecal or intestinal material from infected pigs.7 More recently, sensitive ELISAs in which semi-purified PEDV grown in cell cultures (whole virus), or recombinant proteins or polypeptides of PEDV structural proteins are employed have been developed for the detection of serum antibodies.27 The S1 subunit of the S protein provides the most specific reactions. This is because the whole virus, or N, M or E proteins have known cross-reactivity with TGEV, PRCV and PDCoV.25, 41 The viral neutralization (VN) test in cell cultures is critical to assess virus neutralizing antibodies to PEDV.20 These serological assays have been widely used to monitor prior exposure to the virus and to evaluate the efficacy of vaccines.
Control
Appropriate biosecurity measures should be applied to prevent the introduction of PED onto farms. Present epidemiological knowledge indicates that virus introductions occur mainly through animal and human traffic, and likely by contaminated feed (https://www.aphis.usda.gov/animal_health/animal_dis_spec/swine/downloads/secd_final_report.pdf).
Animals suffering from PED should have free access to water to reduce dehydration which exacerbates the severity of the disease. It is advisable to temporarily withhold feed from fattening pigs while they are in the acute stage of the disease. Measures that delay the entrance of the virus into farrowing units containing new-born piglets may delay the infection until the piglets are older and thereby result in lower mortality.
The artificial transmission of the infection to pregnant sows (feedback method) promotes rapid development of lactogenic immunity and thus shortens the course of the disease on the farm. Such artificial exposure can be accomplished by use of diarrhoeic faeces or the intestinal contents of dead animals, used either fresh or from frozen material.10 However this method may also enhance the spread of other infectious agents throughout the farm.
In Europe, the disease is considered to be of marginal economic importance and therefore does not warrant the development of a vaccine. Mild S INDEL strains currently appear to be in circulation.
Classical PEDV outbreaks in Asia were so severe in the 1980s and 1990s that attempts were made to develop an attenuated viral vaccine. In China, the first CV777-based inactivated and attenuated PEDV vaccines were approved in 1995 and 1998, respectively,67 while in Korea, attenuated vaccines based on classical PEDV strains KPEDV-9 and DR13 were commercialized in 1999 and 2004, respectively.36, 58 A commercial attenuated PEDV vaccine of cell culture adapted classical PEDV P-5V strain (Nisseiken Co. Ltd., Japan) has been administered to sows since 1997 in Japan.56 The use of such vaccines based on classical PEDV strains mostly controlled PED in Asia before the highly virulent non-S INDEL PEDV strains emerged. As demonstrated in the field, the classical PEDV-vaccinated pigs were insufficiently protected from the emerging highly virulent non-S INDEL PEDV infection.62, 68
Since the outbreaks of 2013, the USA has conditionally licensed two PEDV vaccines based on emerging non-S INDEL PEDV strains. The first vaccine (June 2014) was a subunit vaccine. It was developed by HarrisvaccinesTM (now Merck Animal Health) using a replication deficient Venezuelan equine encephalitis (VEE) virus packaging system to express the S protein of an emerging non-S INDEL PEDV strain.17 The second vaccine (September 2015) developed by Zoetis was an inactivated whole-virus (non-S INDEL PEDV) vaccine plus an undefined adjuvant.17 In October 2016, an inactivated vaccine based on non-S INDEL PEDV strain AJ1102 was licensed in China.67 In South Korea, an inactivated vaccine candidate based on non-S INDEL strain KNU-141112 was confirmed protective in sows and their suckling piglets.1 However, the efficacy of these vaccines/vaccine candidates in the field is unproven. Reverse genetics is a technology that can be used to study the function of viral genes and to generate recombinant viruses with defined genetic changes as vaccine candidates. To date, reverse genetics platforms have been generated for both classical and emerging non-S INDEL PEDV strains using different technologies2, 31, 39 and can be used for the future rationale design of safe and effective PEDV vaccines.
References
- BAEK, P.S., CHOI, H.W., LEE, S., YOON, I.J., LEE, Y.J., LEE DU, S., LEE, S. & LEE, C., 2016. Efficacy of an inactivated genotype 2b porcine epidemic diarrhea virus vaccine in neonatal piglets. Veterinary Immunology and Immunopathology, 174, 45-49.
- BEALL, A., YOUNT, B., LIN, C.M., HOU, Y., WANG, Q., SAIF, L. & BARIC, R., 2016. Characterization of a Pathogenic Full-Length cDNA Clone and Transmission Model for Porcine Epidemic Diarrhea Virus Strain PC22A. Molecular Biosystems, 7, e01451-01415.
- BERNASCONI, C., GUSCETTI, F., UTIGER, A., VAN REETH, K., ACKERMANN, M. & POSPISCHIL, A., 1995. Experimental infection of gnotobiotic piglets with a cell culture adapted porcine epidemic diarrhoea virus: Clinical, histopathological and immunohistochemical findings. Proceedings of the Third Congress of the European Society of Veterinary Virology, Interlaken, Switzerland, 542–546.
- BJUSTROM-KRAFT, J., WOODARD, K., GIMENEZ-LIROLA, L., ROTOLO, M., WANG, C., SUN, Y., LASLEY, P., ZHANG, J., BAUM, D., GAUGER, P., MAIN, R. & ZIMMERMAN, J., 2016. Porcine epidemic diarrhea virus (PEDV) detection and antibody response in commercial growing pigs. BMC Veterinary Research, 12, 99.
- BONIOTTI, M.B., PAPETTI, A., LAVAZZA, A., ALBORALI, G., SOZZI, E., CHIAPPONI, C., FACCINI, S., BONILAURI, P., CORDIOLI, P. & MARTHALER, D., 2016. Porcine Epidemic Diarrhea Virus and Discovery of a Recombinant Swine Enteric Coronavirus, Italy. Emerging Infectious Diseases, 22, 83-87.
- BRIDGEN, A., DUARTE, M., TOBLER, K., LAUDE, H. & ACKERMANN, M., 1993. Sequence determination of the nucleocapsid protein gene of the porcine epidemic diarrhoea virus confirms that this virus is a coronavirus related to human coronavirus 229E and porcine transmissible gastroenteritis virus. Journal of General Virology, 74, 1795–1804.
- CALLEBAUT, P., DEBOUCK, P. & PENSAERT, M.B., 1982. Enzyme linked immunosorbent assay for the detection of the coronavirus-like agent and its antibodies in pigs with porcine epidemic diarrhoea. Veterinary Microbiology, 7, 295–306.
- CALLEBAUT, P., PENSAERT, M.B., MIRY, C., HAESEBROUCK, F. & VERGOTE, J., 1986. Prevalence of influenza-Aujeszky, transmissible gastroenteritis and porcine epizootic diarrhoea virus infections in feeder pigs. Proceedings of the Ninth Congress of the International Pig Veterinary Society, Barcelona, 216.
- CARVAJAL, A., LANZA, I., DIEGO, R., RUBIO, P. & CARMENES, P., 1995. Seroprevalence of porcine epidemic diarrhoea virus infection among different types of breeding swine farms in Spain. Preventive Veterinary Medicine, 23, 33–40.
- CHATTHA, K.S., ROTH, J.A. &SAIF, L.J., 2015. Strategies for design and application of enteric viral vaccines. Annual Review of Animal Biosciences, 3, 375-395.
- CHEN, Q., GAUGER, P.C., STAFNE, M.R., THOMAS, J.T., MADSON, D.M., HUANG, H., ZHENG, Y., LI, G. & ZHANG, J., 2016. Pathogenesis comparison between the United States porcine epidemic diarrhea virus prototype and S-INDEL-variant strains in conventional neonatal piglets. Journal of General Virology, 97, 1107-1121.
- CHEN, Q., LI, G., STASKO, J., THOMAS, J.T., STENSLAND, W.R., PILLATZKI, A.E., GAUGER, P.C., SCHWARTZ, K.J., MADSON, D., YOON, K.J., STEVENSON, G.W., BURROUGH, E.R., HARMON, K.M., MAIN, R.G. & ZHANG, J., 2014. Isolation and Characterization of Porcine Epidemic Diarrhea Viruses Associated with the 2013 Disease Outbreak among Swine in the United States. Journal of Clinical Microbiology, 52, 234-243.
- CHOI, J.C., LEE, K.K., PI, J.H., PARK, S.Y., SONG, C.S., CHOI, I.S., LEE, J.B., LEE, D.H. & LEE, S.W., 2014. Comparative genome analysis and molecular epidemiology of the reemerging porcine epidemic diarrhea virus strains isolated in Korea. Infection, genetics and evolution: Journal of Molecular Epidemiology and Evolutionary Genetics in Infectious Diseases, 26, 348-351.
- CHOUDHURY, B., DASTJERDI, A., DOYLE, N., FROSSARD, J.P. & STEINBACH, F., 2016. From the field to the lab - An European view on the global spread of PEDV. Virus Research, 226, 40-49.
- CHUNG, H.C., LEE, J.H., NGUYEN, V.G., HUYNH, T.M.L., LEE, G.E., MOON, H.J., PARK, S.J., KIM, H.K. & PARK, B.K., 2016. New emergence pattern with variant porcine epidemic diarrhea viruses, South Korea, 2012-2015. Virus Research, 226, 14-19.
- CRAWFORD, K., LAGER, K., MILLER, L., OPRIESSNIG, T., GERBER, P. & HESSE, R., 2015. Evaluation of porcine epidemic diarrhea virus transmission and the immune response in growing pigs. Veterinary Research, 46, 49.
- CRAWFORD, K., LAGER, K.M., KULSHRESHTHA, V., MILLER, L.C. & FAABERG, K.S., 2016. Status of vaccines for porcine epidemic diarrhea virus in the United States and Canada. Virus Research, 226, 108-116.
- DEBOUCK, P., CALLEBAUT, P. & PENSAERT, M.B., 1981. The diagnosis of coronavirus-like agent (CVLA) diarrhoea in suckling pigs. Current Topics in Veterinary Medicine and Animal Science, 13, 59–61.
- DEBOUCK, P., PENSAERT, M.B. & COUSSEMENT, W., 1981. The pathogenesis of an enteric infection in pigs experimentally induced by the coronavirus-like agent CV777. Veterinary Microbiology, 6, 157–165.
- DIEL, D.G., LAWSON, S., OKDA, F., SINGREY, A., CLEMENT, T., FERNANDES, M.H., CHRISTOPHER-HENNINGS, J. & NELSON, E.A., 2016. Porcine epidemic diarrhea virus: An overview of current virological and serological diagnostic methods. Virus Research 226, 60-70.
- DIEP, N.V., NORIMINE, J., SUEYOSHI, M., LAN, N.T. & YAMAGUCHI, R., 2017a. Novel Porcine Epidemic Diarrhea Virus (PEDV) Variants with Large Deletions in the Spike (S) Gene Coexist with PEDV Strains Possessing an Intact S Gene in Domestic Pigs in Japan: A New Disease Situation. Public Library of Science (PLoS one), 12, e0170126.
- DIEP, N.V., SUEYOSHI, M., IZZATI, U., FUKE, N., TEH, A.P.P., LAN, N.T. & YAMAGUCHI, R., 2017b. Appearance of US-like porcine epidemic diarrhoea virus (PEDV) strains before US outbreaks and genetic heterogeneity of PEDVs collected in Northern Vietnam during 2012-2015. Transboundary and emerging diseases.
- DUCATELLE, R., COUSSEMENT, W., PENSAERT, M.B., DEBOUCK, P. & HOORENS, J., 1982. Pathology of experimental CV777 coronavirus enteritis in piglets. I. Histological and histochemical study. Veterinary Pathology, 19, 46–56.
- EGBERINK, H.F., EDERVEEN, J., CALLEBAUT, P. & HORZINEK, M.C., 1988. Characterization of the structural proteins of porcine epizootic diarrhoea virus, strain CV777. American Journal of Veterinary Research, 49, 1320–1324.
- GIMENEZ-LIROLA, L.G., ZHANG, J., CARRILLO-AVILA, J.A., CHEN, Q., MAGTOTO, R., POONSUK, K., BAUM, D.H., PINEYRO, P. & ZIMMERMAN, J., 2017. Reactivity of Porcine Epidemic Diarrhea Virus Structural Proteins to Antibodies against Porcine Enteric Coronaviruses: Diagnostic Implications. Journal of Clinical Microbiology, 55, 1426-1436.
- HOFMANN, M. & WYLER, R., 1988. Propagation of the virus of porcine epidemic diarrhoea in cell culture. Journal of Clinical Microbiology, 26, 2235–2239.
- HOFMANN, M. & WYLER, R., 1990. Enzyme-linked immunosorbent assay for the detection of porcine epidemic diarrhea coronavirus antibodies in swine sera. Veterinary Microbiology, 21, 263-273.
- HOU, Y., LIN, C.M., YOKOYAMA, M., YOUNT, B.L., MARTHALER, D., DOUGLAS, A.L., GHIMIRE, S., QIN, Y., BARIC, R.S., SAIF, L.J. & WANG, Q., 2017. Deletion of a 197-Amino-Acid Region in the N-Terminal Domain of Spike Protein Attenuates Porcine Epidemic Diarrhea Virus in Piglets. Journal of Virology, 91.
- HUANG, Y.W., DICKERMAN, A.W., PINEYRO, P., LI, L., FANG, L., KIEHNE, R., OPRIESSNIG, T. & MENG, X.J., 2013. Origin, evolution, and genotyping of emergent porcine epidemic diarrhea virus strains in the United States. Molecular Biosystems, 4.
- HWANG, E.K., KIM, J.H., JEAN, Y.N., BAE, Y.C., YOON, S.S., PARK, C.K., KWEON, C.H., YOON, Y.D. & ACKERMANN, M., 1994. Current occurrence of porcine epidemic diarrhoea in Korea. Rural Development Administration Journal of Agricultural Science, 36, 578-596.
- JENGARN, J., WONGTHIDA, P., WANASEN, N., FRANTZ, P.N., WANITCHANG, A. & JONGKAEWWATTANA, A., 2015. Genetic manipulation of porcine epidemic diarrhea virus (PEDV) recovered from a full length infectious cDNA clone. Journal of General Virology, 96, 2206-2218.
- JUNG, K., ANNAMALAI, T., LU, Z. & SAIF, L.J., 2015. Comparative pathogenesis of US porcine epidemic diarrhea virus (PEDV) strain PC21A in conventional 9-day-old nursing piglets vs. 26-day-old weaned pigs. Veterinary Microbiology, 178, 31-40.
- JUNG, K., WANG, Q., SCHEUER, K.A., LU, Z., ZHANG, Y. & SAIF, L.J., 2014. Pathology of US porcine epidemic diarrhea virus strain PC21A in gnotobiotic pigs. Emerging Infectious Diseases, 20, 662-665.
- KWEON, C.H., KWON, B.J., JUNG, T.S., KEE, Y.J., HUR, D.H., HWANG, E.K., RHEE, J.C. & AN, S.H., 1993. Isolation of porcine epidemic diarrhoea virus (PEDV) in Korea. Korean Journal of Veterinary Research, 33, 249-254.
- KWEON, C.H., KWON, B.J., KANG, Y.B. & AN, S.H., 1994. Cell adaptation of KPEDV-9 and serological survey on porcine epidemic diarrhea virus (PEDV) infection in Korea. Korean Journal of Veterinary Research, 34, 321-326.
- KWEON, C.H., KWON, B.J., LEE, J.G., KWON, G.O. & KANG, Y.B., 1999. Derivation of attenuated porcine epidemic diarrhea virus (PEDV) as vaccine candidate. Vaccine, 17, 2546-2553.
- LANGEL, S.N., PAIM, F.C., LAGER, K.M., VLASOVA, A.N. & SAIF, L.J., 2016. Lactogenic immunity and vaccines for porcine epidemic diarrhea virus (PEDV): Historical and current concepts. Virus Research 226, 93-107.
- LEE, D.U., KWON, T., JE, S.H., YOO, S.J., SEO, S.W., SUNWOO, S.Y. & LYOO, Y.S., 2016. Wild boars harboring porcine epidemic diarrhea virus (PEDV) may play an important role as a PEDV reservoir. Veterinary Microbiology, 192, 90-94.
- LI, C., LI, Z., ZOU, Y., WICHT, O., VAN KUPPEVELD, F.J., ROTTIER, P.J. & BOSCH, B.J., 2013. Manipulation of the porcine epidemic diarrhea virus genome using targeted RNA recombination. Public Library of Science (PLoS one), 8, e69997.
- LIN, C.M., ANNAMALAI, T., LIU, X., GAO, X., LU, Z., EL-THOLOTH, M., HU, H., SAIF, L.J. & WANG, Q., 2015a. Experimental infection of a US spike-insertion deletion porcine epidemic diarrhea virus in conventional nursing piglets and cross-protection to the original US PEDV infection. Veterinary Research, 46, 134.
- LIN, C.M., GAO, X., OKA, T., VLASOVA, A.N., ESSEILI, M.A., WANG, Q. & SAIF, L.J., 2015b. Antigenic relationships among porcine epidemic diarrhea virus and transmissible gastroenteritis virus strains. Journal of Virology, 89, 3332-3342.
- LIN, C.M., SAIF, L.J., MARTHALER, D. & WANG, Q., 2016. Evolution, antigenicity and pathogenicity of global porcine epidemic diarrhea virus strains. Virus Research, 226, 20-39.
- LIN, C.N., CHUNG, W.B., CHANG, S.W., WEN, C.C., LIU, H., CHIEN, C.H. & CHIOU, M.T., 2014. US-Like Strain of Porcine Epidemic Diarrhea Virus Outbreaks in Taiwan, 2013-2014. The Journal of veterinary medical science / the Japanese Society of Veterinary Science, 76, 1297-1299.
- LOHSE, L., KROG, J.S., STRANDBYGAARD, B., RASMUSSEN, T.B., KJAER, J., BELSHAM, G.J. & BOTNER, A., 2017. Experimental Infection of Young Pigs with an Early European Strain of Porcine Epidemic Diarrhoea Virus and a Recent US Strain. Transboundary and Emerging Diseases. 64, 1380-1386.
- OKA, T., SAIF, L.J., MARTHALER, D., ESSEILI, M.A., MEULIA, T., LIN, C.M., VLASOVA, A.N., JUNG, K., ZHANG, Y. & WANG, Q., 2014. Cell culture isolation and sequence analysis of genetically diverse US porcine epidemic diarrhea virus strains including a novel strain with a large deletion in the spike gene. Veterinary Microbiology, 173, 258-269.
- OUYANG, K., SHYU, D.L., DHAKAL, S., HIREMATH, J., BINJAWADAGI, B., LAKSHMANAPPA, Y.S., GUO, R., RANSBURGH, R., BONDRA, K.M., GAUGER, P., ZHANG, J., SPECHT, T., GILBERTIE, A., MINTON, W., FANG, Y. & RENUKARADHYA, G.J., 2015. Evaluation of humoral immune status in porcine epidemic diarrhea virus (PEDV) infected sows under field conditions. Veterinary Research, 46, 140.
- PARK, J.Y., KIM, C.J., SHIN, K.S., KIM, W.Y., KANG, S.Y., PARK, Y.H., HAN, H.J. & PARK, Y.H., 1995. Sequence analysis and cDNA probe hybridization of the nucleocapsid (N) protein gene of transmissible gastroenteritis virus (TGEV) and porcine epidemic diarrhea virus (PEDV). Korean Journal of Veterinary Research 35, 515-530.
- PARK, S.J., SONG, D.S. & PARK, B.K., 2013. Molecular epidemiology and phylogenetic analysis of porcine epidemic diarrhea virus (PEDV) field isolates in Korea. Archives of Virology, 158, 1533-1541.
- PENSAERT, M.B. & DEBOUCK, P., 1978. A new coronavirus-like particle associated with diarrhoea in swine. Archives of Virology, 58, 243-247.
- PENSAERT, M.B., CALLEBAUT, P. & DEBOUCK, P., 1982. Porcine epidemic diarrhoea (PED) caused by a coronavirus: Present knowledge. Proceedings of the Seventh International Congress of the Pig Veterinary Society, Mexico City.
- PENSAERT, M.B. & MARTELLI, P., 2016. Porcine epidemic diarrhea: A retrospect from Europe and matters of debate. Virus Research, 226, 1-6.
- PIJPERS, A., VAN NIEUWSTADT, A.P., TERPSTRA, C. & VERHEIJDEN, J.H., 1993. Porcine epidemic diarrhoea virus as a cause of persistent diarrhoea in a herd of breeding and finishing pigs. The Veterinary Record, 132, 129-131.
- POSPISCHIL, A., HESS, R.G. & BACHMANN, P.A., 1981. Light microscopy and ultrahistology of intestinal changes in pigs infected with enzootic diarrhoea virus (EVD): Comparision with transmissible gastroenteritis (TEG) virus and porcine rotavirus infections. Zentralblatt für Veterinärmedizin Reihe B, 28,564-577.
- PURANAVEJA, S., POOLPERM, P., LERTWATCHARASARAKUL, P., KESDAENGSAKONWUT, S., BOONSOONGNERN, A., URAIRONG, K., KITIKOON, P., CHOOJAI, P., KEDKOVID, R., TEANKUM, K. & THANAWONGNUWECH, R., 2009. Chinese-like strain of porcine epidemic diarrhea virus, Thailand. Emerging Infectious Diseases, 15, 1112-1115.
- SATO, T., TAKEYAMA, N., KATSUMATA, A., TUCHIYA, K., KODAMA, T. & KUSANAGI, K., 2011. Mutations in the spike gene of porcine epidemic diarrhea virus associated with growth adaptation in vitro and attenuation of virulence in vivo. Virus Genes, 43, 72-78.
- SONG, D. & PARK, B., 2012. Porcine epidemic diarrhoea virus: a comprehensive review of molecular epidemiology, diagnosis, and vaccines. Virus Genes, 44, 167-175.
- SONG, D.S., OH, J.S., KANG, B.K., YANG, J.S., MOON, H.J., YOO, H.S., JANG, Y.S. & PARK, B.K., 2007. Oral efficacy of Vero cell attenuated porcine epidemic diarrhea virus DR13 strain. Research in Veterinary Science 82, 134-140.
- STEVENSON, G.W., HOANG, H., SCHWARTZ, K.J., BURROUGH, E.R., SUN, D., MADSON, D., COOPER, V.L., PILLATZKI, A., GAUGER, P., SCHMITT, B.J., KOSTER, L.G., KILLIAN, M.L. & YOON, K.J., 2013. Emergence of Porcine epidemic diarrhea virus in the United States: clinical signs, lesions, and viral genomic sequences. Journal of veterinary diagnostic investigation: official publication of the American Association of Veterinary Laboratory Diagnosticians, Inc, 25, 649-654.
- SUEYOSHI, M., TSUDA, T., YAMAZAKI, K., YOSHIDA, K., NAKAZAWA, M., SATO, K., MINAMI, T., IWASHITA, K., WATANABE, M., SUZUKI, Y. & MORI M., 1995. An immunohistochemical investigation of porcine epidemic diarrhoea. Journal of Comparative Pathology, 113, 59-67.
- SUN, D., WANG, X., WEI, S., CHEN, J. & FENG, L., 2016. Epidemiology and vaccine of porcine epidemic diarrhea virus in China: a mini-review. The Journal of Veterinary Medical Science, 78, 355-363.
- SUN, R.Q., CAI, R.J., CHEN, Y.Q., LIANG, P.S., CHEN, D.K. & SONG, C.X., 2012. Outbreak of porcine epidemic diarrhea in suckling piglets, China. Emerging Infectious Diseases, 18, 161-163.
- SUZUKI, T., MURAKAMI, S., TAKAHASHI, O., KODERA, A., MASUDA, T., ITOH, S., MIYAZAKI, A., OHASHI, S. & TSUTSUI, T., 2015. Molecular characterization of pig epidemic diarrhoea viruses isolated in Japan from 2013 to 2014. Infection, Genetics and Evolution: Journal of Molecular Epidemiology and Evolutionary Genetics in Infectious Diseases, 36, 363-368.
- SUZUKI, T., SHIBAHARA, T., YAMAGUCHI, R., NAKADE, K., YAMAMOTO, T., MIYAZAKI, A. & OHASHI, S., 2016. Pig epidemic diarrhoea virus S gene variant with a large deletion non-lethal to colostrum-deprived newborn piglets. The Journal of General Virology, 97, 1823-1828.
- VAN REETH, K. & PENSAERT, M., 1994. Prevalence of infections with enzootic respiratory and enteric viruses in feeder pigs entering fattening herds. The Veterinary Record, 135, 594-597.
- VLASOVA, A.N., MARTHALER, D., WANG, Q., CULHANE, M.R., ROSSOW, K.D., ROVIRA, A., COLLINS, J. & SAIF, L.J., 2014a. Distinct characteristics and complex evolution of PEDV strains, North America, May 2013-February 2014. Emerging Infectious Diseases, 20, 1620-1628.
- WANG, D., FANG, L. & XIAO, S., 2016. Porcine epidemic diarrhea in China. Virus Research, 226, 7-13.
- WANG, J., ZHAO, P., GUO, L., LIU, Y., DU, Y., REN, S., LI, J., ZHANG, Y., FAN, Y., HUANG, B., LIU, S. & WU, J., 2013. Porcine epidemic diarrhea virus variants with high pathogenicity, China. Emerging Infectious Diseases, 19, 2048-2049.
- WANG, L., BYRUM, B., & ZHANG, Y., 2014. New variant of porcine epidemic diarrhea virus, United States, 2014. Emerging Infectious Diseases, 20, 917-919.
- WITTE, K.H. & PRAGER, D., 1987. Der Nachweis von Antikörpern degren das virus der Epizootischen Virusdiarrhoe (EVD) des Schweines mit dem Immunofloreszenz-blockadetest (IFBT). Tierärztliche Umschau, 42, 817-820.
- WOOD, E.N., 1977. An apparently new syndrome of porcine epidemic diarrhoea. The Veterinary Record, 100, 243-244.
- YALING, R., EDERVEEN, K., EGBERINK, H., PENSAERT, M.B. & HORZINEK, M.C., 1988. Porcine epidemic diarrhoea virus (CV777) and feline infections peritonitis virus (FIPV) are antigenically related. Archives of Virology, 102, 63-71.